The topic for this blog post is how to create a composite gif made up of multiple gganimate objects, overlaid on top of one another.
This is not fundamentally novel, as these abilities are hinted at in the documentation of both gganimate and magick, but to my knowledge this is the first actual demonstration of a multi-panel composite of overlaid gif animations using gganimate and magick.
My original reason for attempting this animation was to make an eye-catching centerpiece for a poster we presented at the ISF 2021 conference, and since the conference was entirely web-based (thanks COVID-19!) and the figure I was basing it on was created with ggplot2, the idea of creating a gif using gganimate was not far-fetched.
The original, static, multi-panel plot
In the work described in our paper published last year (see citation in figure caption) we tracked a chemical reaction of a colloidal solution of zinc oxide nanoparticles (ZnO NPs) in the presence of an common dye (methylene blue, MB) under simulated solar illumination using UV/Vis absorbance spectroscopy.
.](assets/fig-Ahmed-JPhysChemC-2020.png)
Figure 1: Reproduced from J. Phys. Chem. C 2020, 124, 11, 6395.
Since we measured a full spectrum every minute, and the reaction usually ran for over 100 minutes, it was natural to display the data as a time evolution series, showing how the spectral bands change over the course of the reaction.
From the band edge, we deduced the optical band gap of the ZnO NPs, from which we could calculate the average diameter of the NPs. From the MB absorption band we got the absorbance of MB, which decreased over time as the photocatalyst degraded the dye.
So, not the most polished plot ever, but I tend to value technicality over aesthestics most days of the week. The plot was created exclusively using ggplot2, and in so doing I supposed we lived up to the sentiment expressed by Hadley Wickham in this 2020 interview by Will Chase:
I hope that more people continue to express visualizations in a programming language, rather than a point and click tool. That’s my hope for the next 20 years, that it perchance becomes a given that code is the language of data science and data visualization.
The source code generating the static plot is published as part of the git repository of the paper.
I should note, that we use a few technical tricks (for lack of a better word) in this plot, for example, the plotted absorbance values are squared and at the same time the ordinate is shown using a square-root transform. This allows us to visually magnify the MB absorbance bands, which would otherwise be dwarfed by the prominent band edge absorbance. We use a similar trick for the abscissa, by plotting it against the energy scale instead of the wavelength scale, which has the effect of expanding the band edge region horizontally, making it much easier to follow the shift of the band edge over time.
Finally, to make the plot more information-dense (chemists just hate to leave unused whitespace in plots),
we made use of the empty space created in the plot by our stretching and squeezing of the axes
by plotting the derived quantities from the band edge (band gap and particle diameter) and the fitted
absorbance as inset plots (we used grid::viewport() to position the insets).
Now, let’s see how we went about “translating” this into a multi-panel animation.
Animated plot with multiple overlaid panels
Here’s the animated plot:
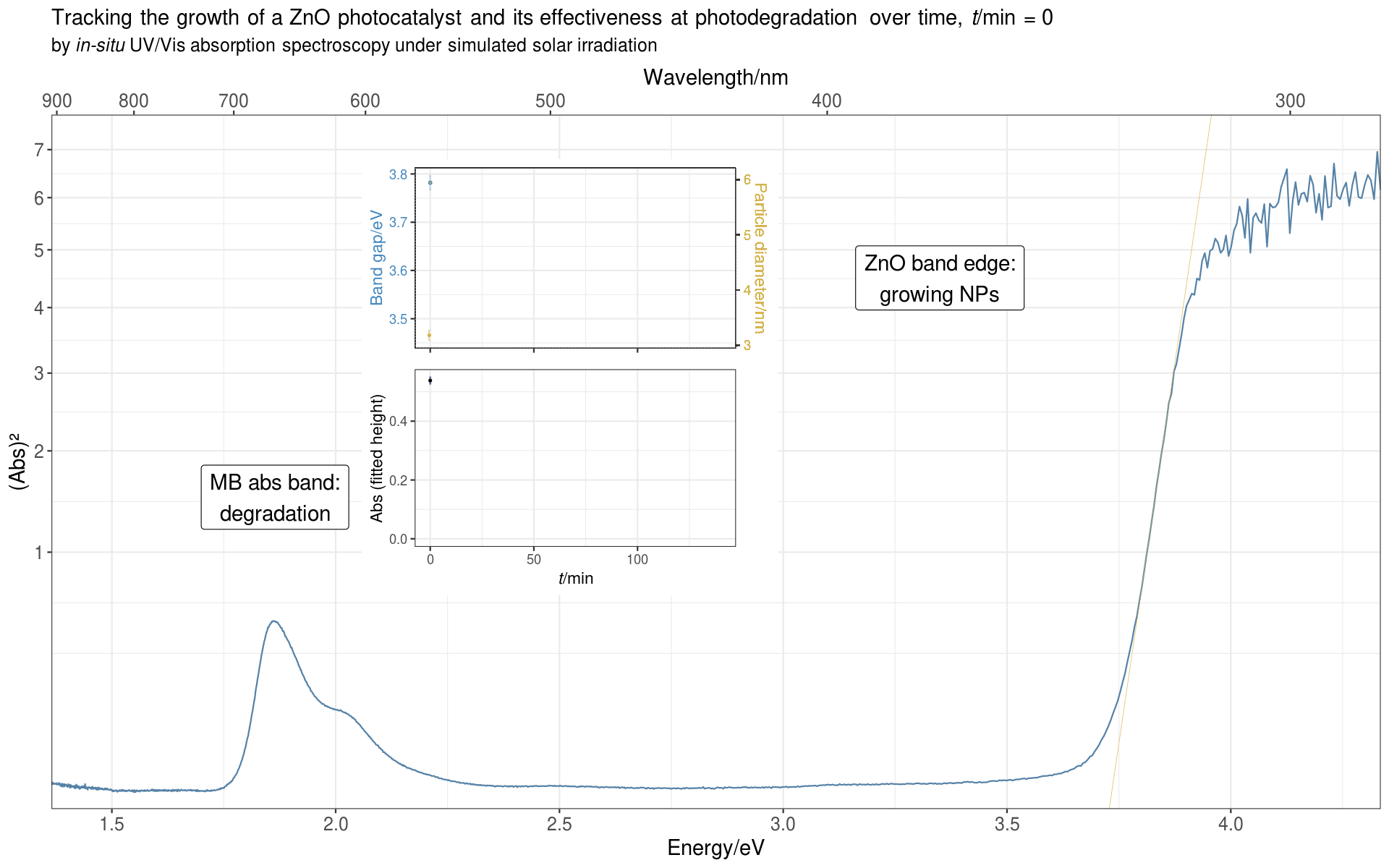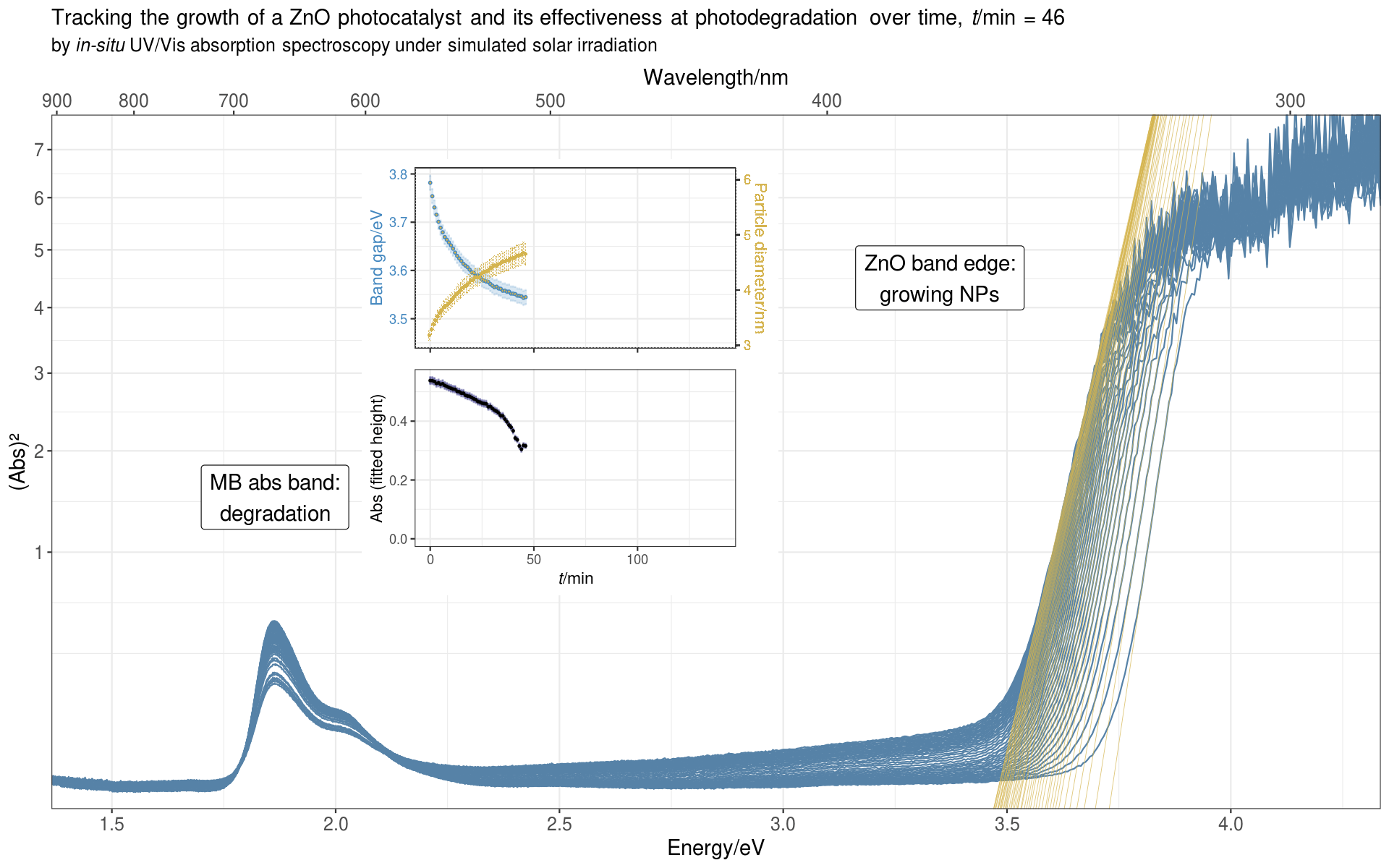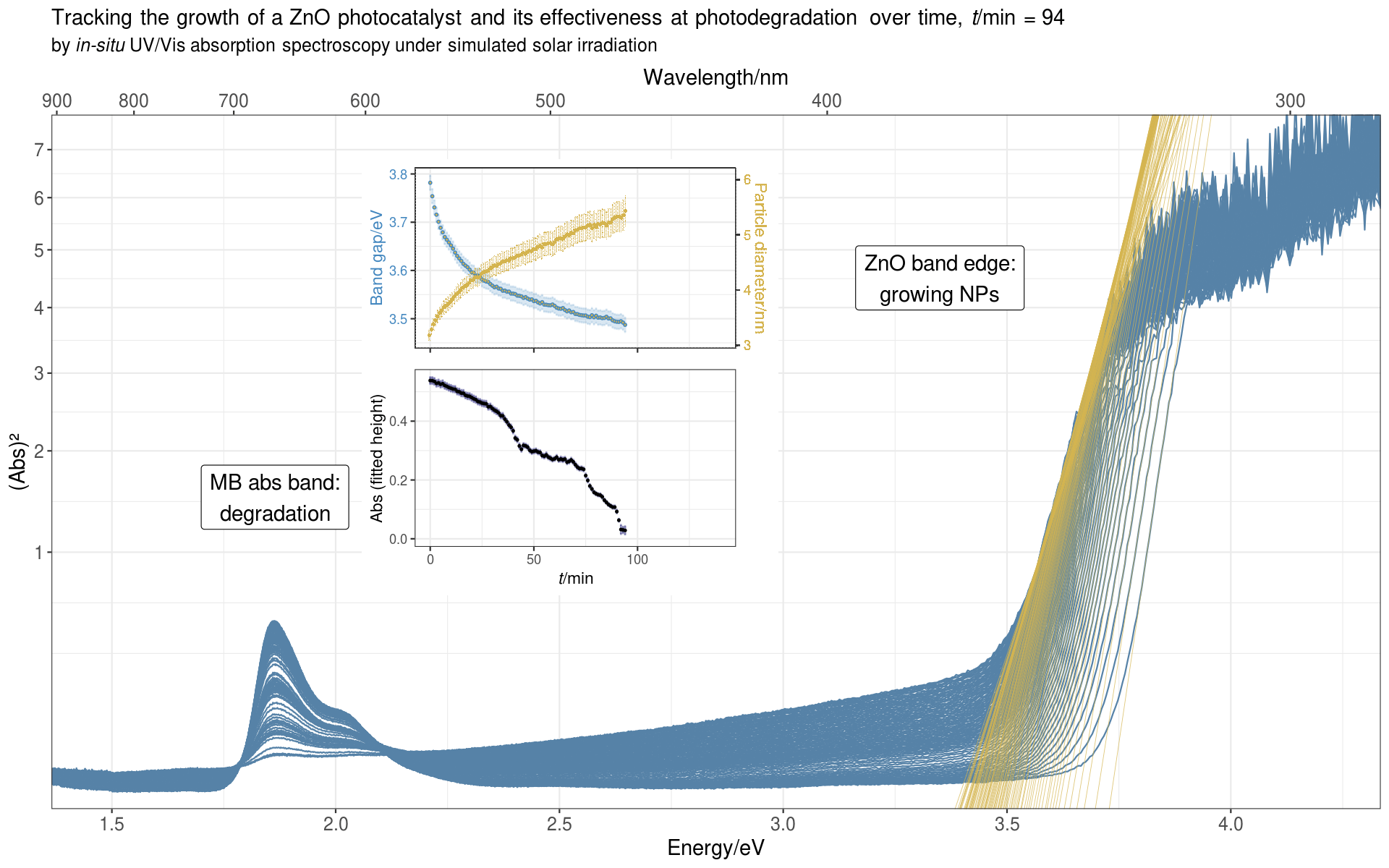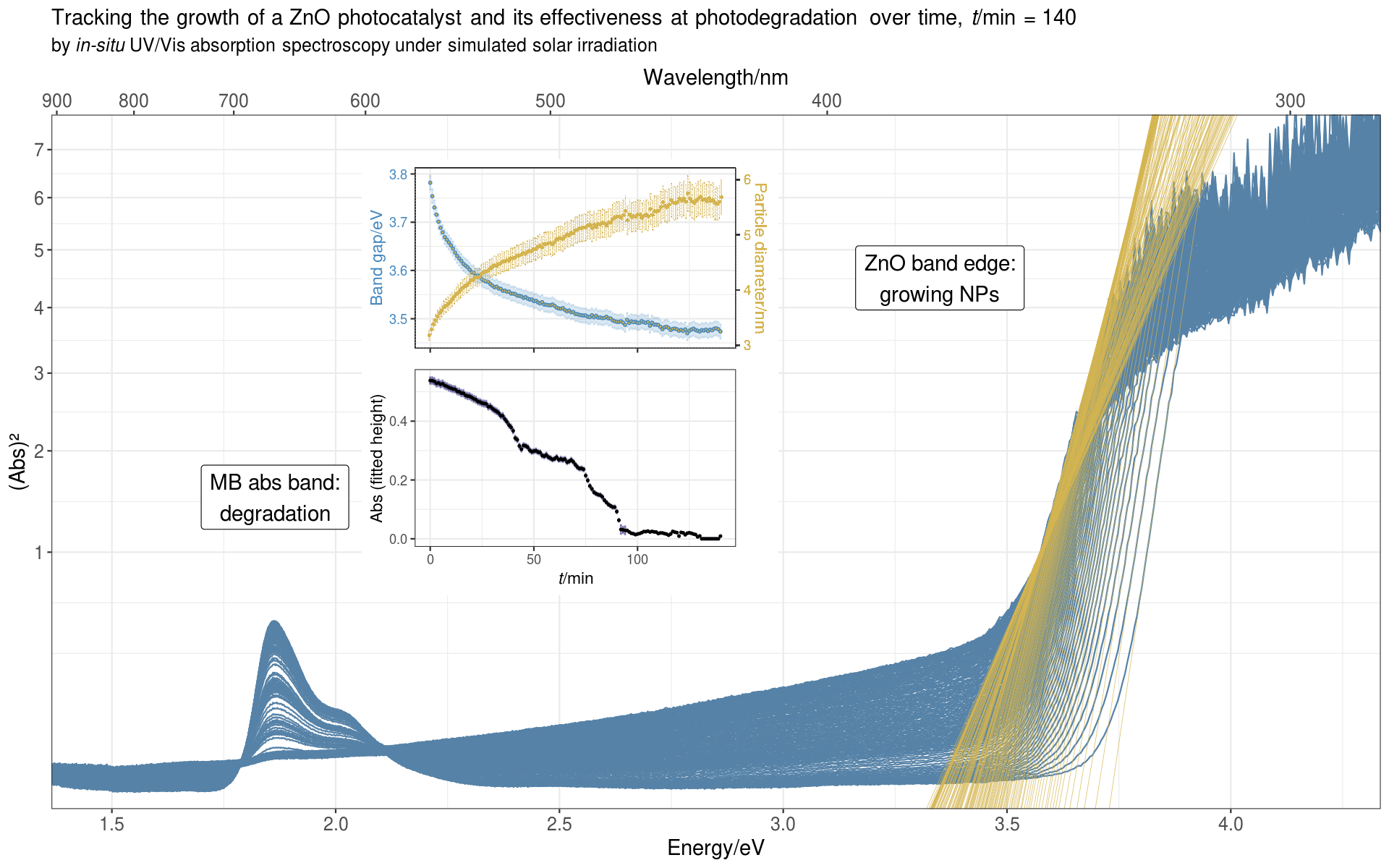
For the intrepid reader who wants to see it all,
the full source code of this blog post is available.
For everyone else, the following chunks demonstrate just the ggplot2::, gganimate::
and magick:: functions we used to build the animated plot above.
nth_spectra <- 1
this_data <-
pc_abs_small_N04H_nostir %>%
filter(time_abs %in%
seq(0, length(unique(pc_abs_small_N04H_nostir$time_abs)),
by = nth_spectra))
this_condensed <- condensed_abs_small_N04H_nostir
this_ceiling <- ceiling_pc_abs_small_N04H_nostir
this_fityk <-
fityk_peaks_small_N04H_nostir_MB_max %>%
# rename to conform with the name used in this_data and this_ceiling
rename(time_exp = spectra)
mb_yoffset <- 220
cg_xoffset <- 500
cg_width <- 576
cg_height <- 324
# this height adjustment makes the plot panels of both insets equally sized
# necessary because only the bottom inset includes x-axis title and labels
mb_height <- cg_height - 45
# note, mb_yoffset actually pertains to the edge/diam plots, and vice versa
edge_yoffset <- mb_yoffset + mb_height
diam_yoffset <- mb_yoffsetFor convenience, I rename the dataframes that will make up the different plots, and define coordinates and dimensions (in pixels) of the inset plots.
p <-
ggplot() +
geom_line(
data = this_data,
colour = "#5682a7",
aes(x = eV, y = sqabs, group = sampleid)) +
## Tauc linear fits
geom_smooth(
data = subset(this_ceiling, fitted == TRUE),
aes(x = eV, y = sqabs, group = time_exp),
method = "lm",
formula = "y ~ x",
se = FALSE,
size = 0.2,
colour = alpha("#D5B450", 0.75),
linetype = 1,
fullrange = TRUE) +
# text box "ZnO band edge"
annotate(
"label",
x = 3.35, y = 4.5,
label = "ZnO band edge:\ngrowing NPs",
size = 5, label.padding = unit(0.4, "lines")) +
# text box "MB abs band"
annotate(
"label",
x = wavelength2energy(MB_band), y = 1.5,
label = "MB abs band:\ndegradation",
size = 5, label.padding = unit(0.4, "lines")) +
# axis labels
labs(
x = "Energy/eV", y = "(Abs)²",
title =
paste(
"Tracking the growth of a ZnO photocatalyst and its effectiveness",
"at photodegradation over time, *t*/min = {round(frame_along, 0)}"),
subtitle =
paste(
"by *in-situ* UV/Vis absorption spectroscopy",
"under simulated solar irradiation")) +
# secondary x-axis (wavelength)
scale_x_continuous(
breaks = seq(1, 5, 0.5),
sec.axis = sec_axis(~ 1239.842 / ., name = "Wavelength/nm")) +
scale_y_sqrt(expand = c(0, 0.05), breaks = seq(0, 8)) +
coord_cartesian(xlim = c(1.5, 4.2), ylim = c(0, 7.5)) +
theme_bw() +
theme(
legend.position = "none",
axis.title = element_text(size = 14),
axis.text = element_text(size = 12),
plot.title = element_markdown(size = 14),
plot.subtitle = element_markdown(size = 12))
# create animated plot
p_anim <- p + transition_reveal(along = time_exp)
p_anim_gif <-
animate(
plot = p_anim,
renderer = gifski_renderer(),
width = 1920, height = 1200, units = "px",
res = 150, # larger res makes all plot elements appear bigger
# one frame per UV/Vis spectrum
nframes = max(this_data$time_exp))It is definitely nice to be able to use Markdown in the title, subtitle and text annotations,
and the functions of the ggtext:: package allows us to do that.
We used the gifski renderer, which is the default, but I think it requires the gifski package to be installed on the system. I created an ansible role to take care of that.
p_mb <-
ggplot(data = this_fityk %>% filter(peakno %% 2 == 1)) +
geom_errorbar(
colour = alpha("#0B096C", 0.4),
size = 0.65,
aes(
x = time_exp,
group = seq_along(time_exp),
ymin = errors_min(height),
ymax = errors_max(height))) +
geom_point(
colour = "black",
size = 0.5,
aes(
x = time_exp,
group = seq_along(time_exp),
y = height)) +
labs(x = "*t*/min", y = "Abs (fitted height)") +
scale_y_continuous(
# easiest way to make certain this panel matches width of diam
# is to give both the same sec_axis
sec.axis =
sec_axis(
trans = ~.,
name = "Particle diameter/nm",
breaks = seq(0, 0.4, 0.2),
labels = c(3,4,5))) +
theme_bw() +
theme(
legend.position = "none",
axis.title.x = element_markdown(),
axis.title.y.right = element_text(colour = NA),
axis.text.y.right = element_text(colour = NA),
axis.ticks.y.right = element_blank(),
# make area outside panel transparent
plot.background = element_rect(fill = NA, colour = NA))
# create animated plot
p_mb_anim <- p_mb + transition_reveal(along = time_exp)
p_mb_anim_gif <-
animate(
plot = p_mb_anim,
renderer = gifski_renderer(),
width = cg_width, height = cg_height, units = "px",
res = 150,
nframes = max(this_data$time_exp))The MB inset plot is pretty straight-forward.
In all the insets, we continue to transition_reveal along the same variable, and we use
the same number of frames as in the main plot.
Note the use of seq_along() in the group aesthestics, which is necessary for the visual
effect of the data points sequentially revealing themselves (with previously shown points
remaining visible).
main_gif <- image_read(p_anim_gif)
mb_gif <- image_read(p_mb_anim_gif)
cg_offset <- paste0("+", cg_xoffset, "+", edge_yoffset)
cg_gravity <- "NorthWest"
main_mb_gif <-
image_composite(
image = main_gif[1],
composite_image = mb_gif[1],
offset = cg_offset,
gravity = cg_gravity)
for (i in 2:max(this_data$time_exp)) {
combined <-
image_composite(
image = main_gif[i],
composite_image = mb_gif[i],
offset = cg_offset,
gravity = cg_gravity)
main_mb_gif <- c(main_mb_gif, combined)
}With both the main plot and the first inset created (as gganimate objects), it is time to create the first composite animation. The loop is a little awkward, and can probably be coded more efficiently, but hey, it works.
This gave us a new gganimate object, main_mb_gif, consisting of the main plot and one inset.
p_edge <-
ggplot(data = this_condensed) +
geom_errorbar(
size = 0.65,
colour = alpha("#4387BF", 0.2),
aes(
x = time_exp,
group = seq_along(time_exp),
ymin = errors_min(fit_Eg),
ymax = errors_max(fit_Eg))) +
geom_point(
shape = 21,
size = 0.65,
fill = "#D5B450",
colour = "#4387BF",
aes(
x = time_exp,
group = seq_along(time_exp),
y = fit_Eg)) +
labs(x = "", y = "Band gap/eV") +
scale_x_continuous() +
scale_y_continuous(
sec.axis =
sec_axis(
trans = ~.,
name = "Particle diameter/nm",
breaks = seq(3.5, 3.8, 0.1),
labels = c(3,4,5,6))) +
theme_bw() +
theme(
legend.position = "none",
axis.title.x = element_blank(),
axis.text.x = element_blank(),
axis.title.y = element_text(colour = "#4387BF"),
axis.text.y = element_text(colour = "#4387BF"),
axis.title.y.right = element_text(colour = NA),
axis.text.y.right = element_text(colour = NA),
axis.ticks.y.right = element_blank(),
# make area outside panel transparent
plot.background = element_rect(fill = NA, colour = NA))
# create animated plot
p_edge_anim <- p_edge + transition_reveal(along = time_exp)
p_edge_anim_gif <-
animate(
plot = p_edge_anim,
renderer = gifski_renderer(),
width = cg_width, height = mb_height, units = "px",
res = 150,
nframes = max(this_data$time_exp))
edge_gif <- image_read(p_edge_anim_gif)For the bottom one of the two stacked insets (p_edge and p_diam will be drawn on top of each other),
we need to start getting a little messy with our code.
In order to precisely match the dimensions of the two stacked plots,
and since the bottom one will have a left y-axis whereas the top one needs a right-side y-axis,
both plots get dual y-axes, where the bottom one is drawn invisibly by setting its theme
elements to colour=NA.
We just need to make sure that both y-axes of both plots occupy the same amount of horizontal space,
which we do by setting the labels of each so they are identical.
cg_offset <- paste0("+", cg_xoffset, "+", mb_yoffset)
cg_gravity <- "NorthWest"
main_mb_edge_gif <-
image_composite(
image = main_mb_gif[1],
composite_image = edge_gif[1],
offset = cg_offset,
gravity = cg_gravity)
for (i in 2:max(this_data$time_exp)) {
combined <-
image_composite(
image = main_mb_gif[i],
composite_image = edge_gif[i],
offset = cg_offset,
gravity = cg_gravity)
main_mb_edge_gif <- c(main_mb_edge_gif, combined)
}Now we simply composite the first composite gif (which consists of the main plot with one inset) with the new inset, resulting in a new composite consisting of the main plot with two insets.
p_diam <-
ggplot(
data =
this_condensed %>%
filter(spectra <= subset(cutoffs, sample == "N04H-small-nostir")$diameter)) +
geom_errorbar(
size = 0.35,
colour = alpha("#D5B450", 0.3),
aes(
x = time_exp,
group = seq_along(time_exp),
ymin = errors_min(fit_np_diam),
ymax = errors_max(fit_np_diam))) +
geom_point(
size = 0.45,
colour = "#D5B450",
aes(
x = time_exp,
group = seq_along(time_exp),
y = fit_np_diam)) +
labs(y = "Band gap/eV", x = "") +
# manual adjustments to make sure the diam panel matches the edge panel's size
scale_y_continuous(
# invisible labels on left-hand y-axis, just to occupy space
breaks = seq(3,6), labels = c("3.5", "3.6", "3.7", "3.8"),
sec.axis = sec_axis(~., name = "Particle diameter/nm")) +
scale_x_continuous() +
coord_cartesian(xlim = c(0, max(this_condensed$time_exp))) +
theme_bw() +
theme(
legend.position = "none",
panel.grid = element_blank(),
axis.title.y = element_text(colour = NA),
axis.text.y = element_text(colour = NA),
axis.ticks.y.left = element_blank(),
axis.title.y.right = element_text(colour = "#D5B450"),
axis.text.y.right = element_text(colour = "#D5B450"),
axis.title.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.x = element_blank(),
# make plot background transparent
panel.background = element_rect(fill = NA, colour = NA),
# make area outside panel transparent
plot.background = element_rect(fill = NA, colour = NA))
# create animated plot
p_diam_anim <- p_diam + transition_reveal(along = time_exp)
p_diam_anim_gif <-
animate(
plot = p_diam_anim,
renderer = gifski_renderer(),
bg = "transparent", # https://github.com/thomasp85/gganimate/issues/174
width = cg_width, height = mb_height, units = "px",
res = 150,
nframes = max(this_data$time_exp))
diam_gif <- image_read(p_diam_anim_gif)Here, for the top inset, we re-use the same techniques as for the bottom inset,
with the addition that we need to make sure both panel and plot background is transparent.
Also, we need to make the gganimate transparent, which is achieved with the bg='transparent'
option.
cg_offset <- paste0("+", cg_xoffset, "+", diam_yoffset)
cg_gravity <- "NorthWest"
main_mb_edge_diam_gif <-
image_composite(
image = main_mb_edge_gif[1],
composite_image = diam_gif[1],
offset = cg_offset,
gravity = cg_gravity)
for (i in 2:max(this_data$time_exp)) {
combined <-
image_composite(
image = main_mb_edge_gif[i],
composite_image = diam_gif[i],
offset = cg_offset,
gravity = cg_gravity)
main_mb_edge_diam_gif <- c(main_mb_edge_diam_gif, combined)
}Finally, we composite again.
image_write_gif(
image = main_mb_edge_diam_gif,
loop = 10,
path = here::here("assets/animation-composite.gif"))The final composite is then saved as a gif file using magick::image_write_gif().
At this point, we also set loop= to some number so that the gif does not loop forever (just a matter of taste).
Encountered issues and how to handle them
- If you use
knitrtogether withgganimate, be very mindful about knitr’s chunk optiondev! I usually setopts_chunk$set(dev='svg')globally in my blog posts, and it turns outgganimate::animate(device=..)inherits this chunk option. Obviously the SVG device makes no sense for a gif, and you will get not-very-useful device-related errors. So if you are using knitr, make sure to setdevice='png'inside each call toanimate(), or else in the relevant chunk’s options. magick::image_read()failed with the errorunable to get registry ID cache:hosts @ error/registry.c/GetImageRegistry/202for no apparent reason. Fixed by increasing ImageMagick’s resource limits.magick::image_write_gif()failed with the same error, despiteimage_read()working. Fixed by increasing the resource limits even more (see my imagemagick ansible role).- The
end_pauseargument ofgganimate::animate()has no effect for gifs composited withmagick::image_write_gif(). And unfortunately,image_write_gif()has no similar argument. I thought I could create and end pause manually by repeating the last frame of the gif a number of times, but I could not figure out how to achieve that using the magick functions. In the end, I gave up on adding an end pause and used theloopargument instead to set a max number of loops, since my point was to allow the reader some time to read the plot without everything moving around.
Further reading
gganimate
Creating animations with gganimate
- Building an animation step-by-step with gganimate: mapping the Japanese cherry blossom front, Alex Cookson. 2020-10-19.
- gganimate (with a spooky twist), Katherine Goode. ISU Graphics group, 2019-10-31.
- Some fun with {gganimate}, Bruno Rodrigues. Econometrics and Free Software, 2018-12-27.
- Working with images in R, Jeroen Ooms. Institute for Geoinformatics, WWU Münster, 2018-10-16.
- gganimate: animations with ggplot2, Thean Lim. 2018-04-19.
- Iteration and animation: loops, GIFs, and videos, Peter Aldhous. 2018.
- How to create animations in R with gganimate, Ander Fernández Jauregui.
gganimate::animate()documentation.
Composite animations with gganimate
- The magick package: advanced image-processing in R, magick package vignette. 2021-07-30.
- Multiple gganimate plots both stacked and side by side, StackOverflow question by Abdel, answer by user12728748. 2020-05-20.
- Combining two animated plots into one GIF/MP4, StackOverflow question by Keith McNulty, answer by Stéphane Laurent. 2019-05-03.
- Placing animations side-by-side with magick, Matt Crump. gganimate wiki, 2018-08-17.
magick::image_composite()documentation.
Composite animations in R without using gganimate
sessionInfo()
## R version 4.0.5 (2021-03-31)
## Platform: x86_64-pc-linux-gnu (64-bit)
## Running under: Ubuntu 18.04.6 LTS
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.7.1
## LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.7.1
##
## locale:
## [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
## [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
## [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
## [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] conflicted_1.1.0 photoec_0.2.2.9000 oceanoptics_0.0.0.9004
## [4] common_0.0.2 tibble_3.1.6 tidyr_1.1.4
## [7] dplyr_1.0.8 magrittr_2.0.2 errors_0.3.6
## [10] knitr_1.37 here_1.0.1 ggtext_0.1.1
## [13] magick_2.7.3 gganimate_1.0.7 ggplot2_3.3.5
##
## loaded via a namespace (and not attached):
## [1] progress_1.2.2 tidyselect_1.1.2 xfun_0.29 bslib_0.3.1
## [5] purrr_0.3.4 colorspace_2.0-2 vctrs_0.3.8 generics_0.1.2
## [9] htmltools_0.5.2 yaml_2.2.2 utf8_1.2.2 rlang_1.0.1
## [13] gridtext_0.1.4 jquerylib_0.1.4 pillar_1.7.0 glue_1.6.1
## [17] withr_2.4.3 DBI_1.1.2 tweenr_1.0.2 lifecycle_1.0.1
## [21] stringr_1.4.0 munsell_0.5.0 blogdown_1.7 gtable_0.3.0
## [25] memoise_2.0.1 evaluate_0.14 fastmap_1.1.0 fansi_1.0.2
## [29] gifski_1.4.3-1 highr_0.9 Rcpp_1.0.8 scales_1.1.1
## [33] cachem_1.0.6 jsonlite_1.7.3 farver_2.1.0 png_0.1-7
## [37] hms_1.1.1 digest_0.6.29 stringi_1.7.6 bookdown_0.24
## [41] rprojroot_2.0.2 grid_4.0.5 cli_3.2.0 tools_4.0.5
## [45] sass_0.4.0 crayon_1.5.0 pkgconfig_2.0.3 ellipsis_0.3.2
## [49] xml2_1.3.3 prettyunits_1.1.1 assertthat_0.2.1 rmarkdown_2.11
## [53] R6_2.5.1 compiler_4.0.5